The Science Behind Human Metapneumovirus (hMPV): Molecular Insights and Evolutionary Adaptations
Comprehensive Overview of Human Metapneumovirus (hMPV)
Human metapneumovirus is a non-segmented, enveloped, negative-stranded RNA virus classified under the family Paramyxoviridae, subfamily Pneumovirinae. Ever since its discovery in 2001 in the Netherlands, hMPV has become recognized as one of the major causative agents in acute respiratory tract infections worldwide among young children, the elderly, and people with immunodeficiency.
Classification and Genetic Structure
- Taxonomy: hMPV is closely related to the avian metapneumovirus (APV), forming a separate genus within Pneumovirinae. It lacks hemagglutinin-neuraminidase activity, differentiating it from other members of its family.
- Genome: The RNA genome of hMPV (~13 kb) encodes nine proteins:
- Nucleoprotein (N): Encapsulates the viral genome.
- Phosphoprotein (P): Plays a role in RNA synthesis.
- Matrix protein (M): Involved in virion assembly.
- Fusion protein (F): Essential for viral entry into host cells and a target for neutralizing antibodies.
- M2-1 and M2-2 proteins: Regulate transcription and replication.
- Small hydrophobic protein (SH): Function remains uncertain.
- Attachment protein (G): Mediates viral attachment to host cells.
- Large polymerase protein (L): Catalyzes RNA synthesis.

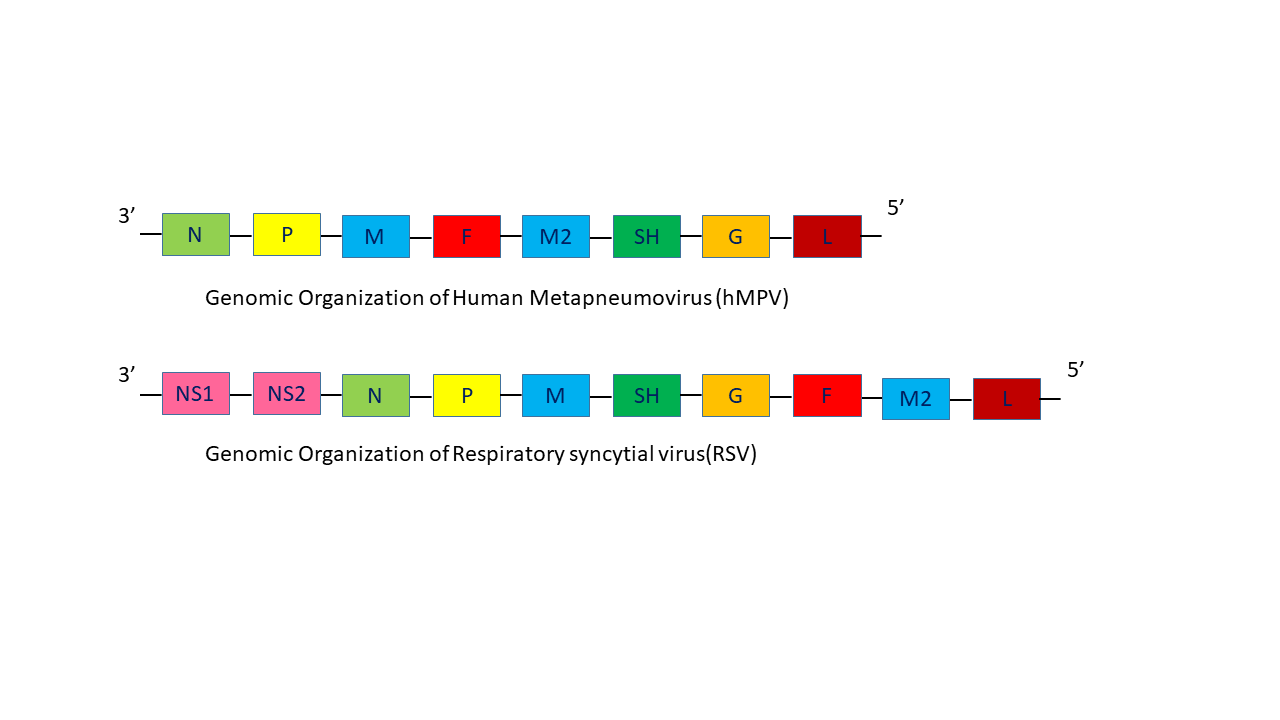
- Phylogenetics: hMPV is classified into two major genetic lineages (A and B), each subdivided into two subgroups (A1, A2, B1, B2). Subgroup A2 is further split into A2a, A2b, and A2c based on variability in the G and F proteins. These lineages co-circulate globally, with varying dominance across regions and seasons.
Epidemiology
- Global Prevalence: hMPV is ubiquitous, infecting nearly all individuals by age five. Its prevalence among Acute Respiratory Tract Infections (ARTIs ) varies by region:
- China: Prevalence ranges from 15.88% in Chongqing to <3% in Shanghai.
- India: Responsible for up to 10% of pediatric ARTIs.
- United States: A leading cause of lower respiratory tract infections (LRTIs) in young children, second only to respiratory syncytial virus (RSV).
- Seasonality: hMPV infections typically occur during late winter and spring, aligning with RSV and influenza outbreaks.
- Age Groups at Risk:
- Infants and young children (<2 years): Most severe cases.
- Elderly: Often experience severe complications.
- Immunocompromised individuals: Prolonged viral shedding and higher mortality.
Transmission
hMPV spreads via respiratory droplets, direct contact with infected secretions, and fomites. The virus remains infectious on surfaces for several hours and in respiratory secretions for up to a week after symptom onset.
Clinical Features
- Symptoms: hMPV causes a wide spectrum of illnesses, ranging from mild upper respiratory tract infections (URTIs) to severe LRTIs, including bronchiolitis, pneumonia, and exacerbations of asthma and chronic obstructive pulmonary disease (COPD). Common symptoms include:
- Fever
- Cough
- Nasal congestion
- Wheezing
- Respiratory distress
- Complications: Severe cases may require hospitalization, mechanical ventilation, and intensive care, particularly in high-risk populations.
Diagnosis
- Reverse Transcription Polymerase Chain Reaction (RT-PCR): The gold standard for hMPV detection, targeting conserved regions of the F or N genes.
- Other Diagnostic Methods:
- Viral Culture: Although definitive, it is time-consuming and requires specialized facilities.
- Serology: Detects specific IgM or IgG antibodies but is less useful for acute diagnosis.
- Rapid Antigen Tests: Emerging technologies offer potential for point-of-care testing but lack sensitivity.
Treatment and Prevention
- Current Treatments:
-
- Supportive Care: Oxygen therapy, hydration, and fever management are the mainstays of treatment.
- Antiviral Agents:
- Ribavirin: Broad-spectrum antiviral that inhibits viral RNA polymerase but is limited by side effects and teratogenicity.
- Monoclonal Antibodies: mAb 338 targets the F protein, showing promise in neutralizing all hMPV subgroups in animal models.
- Immunoglobulin Therapy: Administered to immunocompromised patients to reduce disease severity.
-
Preventive Strategies:
- Infection Control: Hand hygiene, surface disinfection, and isolation of infected individuals are crucial.
- Vaccines: No licensed vaccine exists yet, though F protein-based candidates are under development due to its conserved nature across hMPV subgroups.

Viral Characteristics
- Replication: hMPV replicates in tertiary monkey kidney cells and LLC-MK2 cells but shows poor growth in other cell lines (e.g., Vero, A549). Unlike RSV, it does not induce hemagglutination.
- Genetic Variation: The G protein exhibits high sequence variability, while the F protein is relatively conserved, making it a primary target for therapeutics and vaccine development.
- Co-Infections: hMPV frequently co-occurs with other respiratory pathogens, such as RSV, influenza, and bacterial agents, complicating diagnosis and management.
Research Highlights
- Discovery and Evolution: Serological evidence suggests that hMPV has been circulating in humans since the 1950s.
- PERCH Study (2024): Identified hMPV as the second leading cause of severe pneumonia in children under five, often in conjunction with bacterial infections.
- Ongoing Challenges:
- Developing rapid, affordable diagnostic tests.
- Addressing the lack of specific antiviral therapies and vaccines.
My Analysis and Insights:
In-Depth Analysis of G Protein Mutations in hMPV: Phylogenetic, MSA, and Structural Insights
This study analyzed the evolutionary patterns, mutational variations, and structural implications of the G protein in Human Metapneumovirus (HMPV) strains collected from diverse geographical regions and time periods. A combination of phylogenetic, multiple sequence alignment (MSA), and structural analyses provided insights into the adaptive mechanisms and potential functional consequences of observed mutations.
Phylogenetic and Sequence Analysis
- Phylogenetic Insights:
- HMPV strains clustered based on geographic and temporal proximity, indicating distinct evolutionary lineages and regional adaptation.
- Basal nodes, such as the Netherlands2002 and Argentina2002 strains, suggest early divergence in the virus’s evolutionary history.
- Recent strains, including USA2020 and Russia2023, exhibited close genetic relationships with short branch lengths, indicative of fewer accumulated mutations and recent evolutionary events.
- The divergence of China2023, clustering with older strains like Australia2004, suggests evolutionary conservation or potential reintroduction of older lineages into the population.
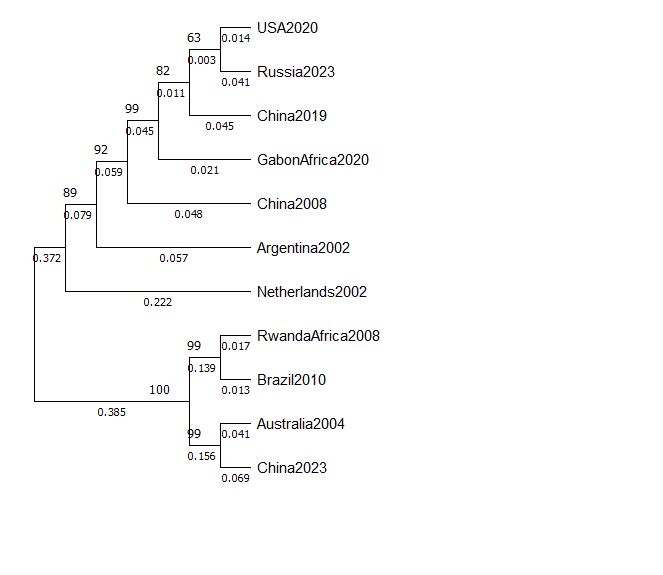
- MSA Highlights:
- Conserved Regions: These regions likely play a critical role in maintaining G protein structure and function, particularly in receptor binding or immune recognition.
- Mutational Hotspots: Significant amino acid substitutions and occasional insertions/deletions were identified in variable regions, which could contribute to strain-specific adaptations.
- Functional Implications: Mutations in conserved domains may enhance receptor affinity or immune evasion, while those in variable regions may reflect localized adaptations to host immune pressures.

Structural Analysis of the G Protein
The structural superimposition of the G protein from different strains revealed minimal overall changes, with a root mean square deviation (RMSD) of 0.001 Å, suggesting high structural conservation. However, a detailed analysis of specific regions provided the following insights:

- Impact of Mutations on Structure:
- Localized Variations: While the global structure remained conserved, mutations in critical regions, such as the receptor-binding domain, may induce subtle side-chain rearrangements that influence interaction dynamics.
- Functional Residues: Changes in conserved residues potentially enhance receptor binding or facilitate immune escape mechanisms, contributing to virulence and transmissibility.
- Stability and Flexibility:
- Conserved structural integrity supports the hypothesis that G protein’s core functionality is preserved despite ongoing mutations.
- Mutations in variable regions may alter local flexibility, which could affect immune recognition or adaptation to host environments.
Integrated Findings
The combination of phylogenetic, sequence, and structural analyses suggests that the G protein of HMPV is undergoing evolutionary adaptation, characterized by regional and temporal divergence. Structural conservation likely ensures the preservation of essential functions such as receptor binding, while mutations in variable regions may drive strain-specific adaptations. The clustering of certain strains with older lineages further highlights the dynamic interplay of evolutionary conservation and adaptation.
Conclusion and Future Directions
The observed genetic and structural variability of the G protein highlights its pivotal role in the evolution and adaptation of HMPV. Specific mutations, particularly in conserved regions, may enhance virulence and immune evasion, potentially influencing infection severity and transmissibility. Continued surveillance of G protein mutations and functional studies are essential to:
- Assess their impact on viral pathogenicity and immune recognition.
- Inform vaccine design and therapeutic interventions.
Understanding the relationship between genetic variability, structural changes, and functional implications will provide a deeper understanding of HMPV evolution and its potential public health impact.
All in all, Human metapneumovirus is a significant global pathogen, causing substantial morbidity and mortality in vulnerable populations. Advances in diagnostics, therapeutics, and vaccines are essential to mitigate its impact and improve patient outcomes. Ongoing research into the molecular and immunological aspects of hMPV holds promise for the development of targeted interventions.
Note: As, hMPV lacks the non structure proteins (NS): the Pathogenicity is quite lower than that of RSV and similar other Viruses. I have only compared the Complete CDS region of G protein that is higly variable compared to others. Lack of Proper sequences among various subtypes limited the study. However, several published research work has reported all the subtypes around the world especially A2c in the Asian regions.
References
- Feuillet, F., Lina, B., Rosa-Calatrava, M., & Boivin, G. (2012). Ten years of human metapneumovirus research. Journal of Clinical Virology, 53(2), 97–105. https://doi.org/10.1016/j.jcv.
- Hamelin, M. E., Abed, Y., & Boivin, G. (2004). Human metapneumovirus: A new player among respiratory viruses. Clinical Infectious Diseases, 38(7), 983–990. https://doi.org/10.1086/382536.
- Costa-Filho, R. C., Saddy, F., Costa, J. L. F., Tavares, L. R., & Castro Faria Neto, H. C. (2025). The silent threat of human metapneumovirus: Clinical challenges and diagnostic insights from a severe pneumonia case. Microorganisms, 13(1), 73. https://doi.org/10.3390/microorganisms13010073.
- Veronese, A., Uršič, T., Bizjak Vojinovič, S., & Rodman Berlot, J. (2024). Exploring clinical predictors of severe human metapneumovirus respiratory tract infections in children: Insights from a recent outbreak. Microorganisms, 12(4), 641. https://doi.org/10.3390/microorganisms12040641.
- P, P., Shetty, U., Parida, P., Varamballi, P., Mukhopadhyay, C., & N, S. (2024). Molecular detection and genotyping of HMPV in patients with severe acute respiratory infection in India. Annals of Medicine, 56(1), 2398719. https://doi.org/10.1080/07853890.2024.2398719.
About the Author:
https://nepomicsbiotech.com.np/member/mr-suman-dhungel/
You can submit your feedback: ∇